
Welcome to PaddleHelix Helper¶


- PaddleHelix is a machine-learning-based bio-computing framework aiming at facilitating the development of the following areas:
Vaccine design
Drug discovery
Precision medicine
Features¶
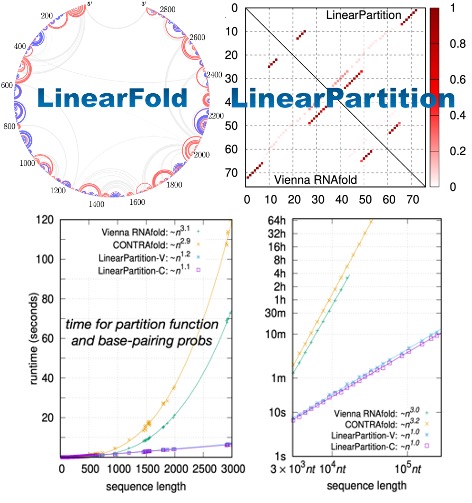
High Efficency: We provide LinearRNA, a highly efficient toolkit for RNA structure prediction and analysis. LinearFold & LinearParitition achieves O(n) complexity in RNA-folding prediction, which is hundreds of times faster than traditional folding techniques.
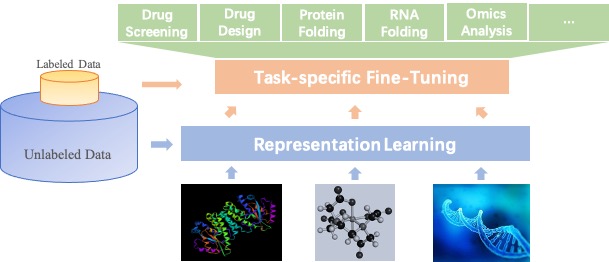
Large-scale Representation Learning and Transfer Learning: Self-supervised learning for molecule representations offers prospects of a breakthrough in tasks with limited annotation, including drug profiling, drug-target interaction, protein-protein interaction, RNA-RNA interaction, protein folding, RNA folding, and molecule design. PaddleHelix implements a variety of representation learning algorithms and state-of-the-art large-scale pre-trained models to help developers to start from “the shoulders of giants” quickly.
Easy-to-use APIs: PaddleHelix provide frequently used structures and pre-trained models. You can easily use those components to build up your models and systems.
Installation¶
OS support¶
Windows, Linux and OSX
Python version¶
Python 3.6, 3.7
Dependencies¶
PaddlePaddle >= 2.0.0rc0
pgl >= 1.2.0
Quick Start¶
PaddleHelix can be installed directly with
pip:
$ pip install paddlehelix
or install from source:
$ pip install --upgrade git+https://github.com/PaddlePaddle/PaddleHelix.git
Note
Please check our Installation part for full installation prerequisites and guide.
Tutorials¶
We provide abundant tutorials to navigate the directory and start quickly.
PaddleHelix is based on PaddlePaddle, a high-performance Parallelized Deep Learning Platform.
Examples¶
Developer’s guide for advanced users¶
If you need help in modifying the source code of PaddleHelix, please see our Developer’s Guide.
Contribution¶
If you would like to develop and maintain PaddleHelix with us, please refer to our GitHub repo.
Overview
Datasets